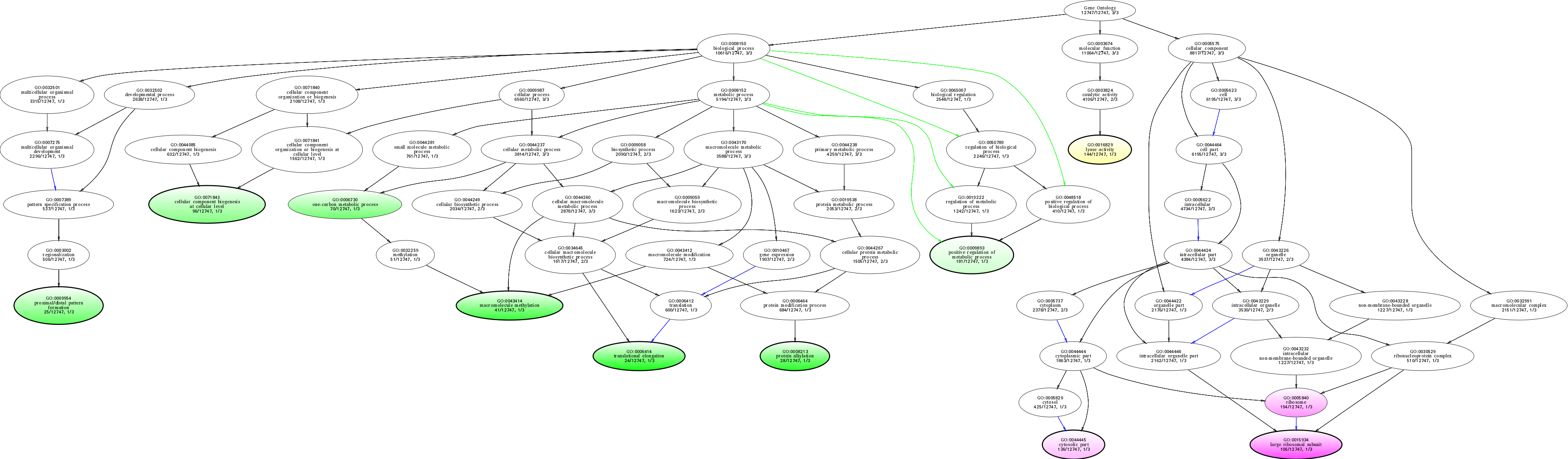
| ID | Name | p-Value | p-Value (Adj) | Study Count | Population Count |
|---|
|
GO:0006414
|
translational elongation
|
0.0294734 |
0.0294734 |
1
|
24
|
|
GO:0008213
|
protein alkylation
|
0.0409357 |
0.0409357 |
1
|
28
|
|
GO:0043414
|
macromolecule methylation
|
0.0420026 |
0.0420026 |
1
|
41
|
|
GO:0015934
|
large ribosomal subunit
|
0.0477693 |
0.0477693 |
1
|
106
|
|
GO:0009954
|
proximal/distal pattern formation
|
0.0494071 |
0.0494071 |
1
|
25
|
|
GO:0006730
|
one-carbon metabolic process
|
0.0536967 |
0.0536967 |
1
|
70
|
|
GO:0071843
|
cellular component biogenesis at cellular level
|
0.0612636 |
0.0612636 |
1
|
96
|
|
GO:0005840
|
ribosome
|
0.0681419 |
0.0681419 |
1
|
194
|
|
GO:0016829
|
lyase activity
|
0.0689196 |
0.0689196 |
1
|
144
|
|
GO:0044445
|
cytosolic part
|
0.0730005 |
0.0730005 |
1
|
136
|
|
GO:0009893
|
positive regulation of metabolic process
|
0.0989758 |
0.0989758 |
1
|
181
|
|
GO:0001071
|
nucleic acid binding transcription factor activity
|
0.103335 |
0.103335 |
1
|
395
|
|
GO:0016741
|
transferase activity, transferring one-carbon groups
|
0.107767 |
0.107767 |
1
|
111
|
|
GO:0051173
|
positive regulation of nitrogen compound metabolic process
|
0.108443 |
0.108443 |
1
|
119
|
|
GO:0048736
|
appendage development
|
0.110896 |
0.110896 |
1
|
286
|
|
GO:0048518
|
positive regulation of biological process
|
0.111456 |
0.111456 |
1
|
410
|
|
GO:0007478
|
leg disc morphogenesis
|
0.111782 |
0.111782 |
1
|
37
|
|
GO:0008152
|
metabolic process
|
0.117083 |
0.117083 |
3
|
5194
|
|
GO:0044237
|
cellular metabolic process
|
0.118315 |
0.118315 |
3
|
3814
|
|
GO:0005198
|
structural molecule activity
|
0.119376 |
0.119376 |
1
|
459
|
|
GO:0010604
|
positive regulation of macromolecule metabolic process
|
0.119673 |
0.119673 |
1
|
150
|
|
GO:0035218
|
leg disc development
|
0.122056 |
0.122056 |
1
|
57
|
|
GO:0010628
|
positive regulation of gene expression
|
0.123263 |
0.123263 |
1
|
122
|
|
GO:0045935
|
positive regulation of nucleobase, nucleoside, nucleotide and nucleic acid metabolic process
|
0.125793 |
0.125793 |
1
|
119
|
|
GO:0031325
|
positive regulation of cellular metabolic process
|
0.125972 |
0.125972 |
1
|
172
|
|
GO:0045941
|
positive regulation of transcription
|
0.127636 |
0.127636 |
1
|
115
|
|
GO:0035127
|
post-embryonic limb morphogenesis
|
0.128205 |
0.128205 |
1
|
35
|
|
GO:0003906
|
DNA-(apurinic or apyrimidinic site) lyase activity
|
0.129630 |
0.129630 |
1
|
7
|
|
GO:0035109
|
imaginal disc-derived limb morphogenesis
|
0.132143 |
0.132143 |
1
|
37
|
|
GO:0007447
|
imaginal disc pattern formation
|
0.133075 |
0.133075 |
1
|
103
|
|
GO:0005811
|
lipid particle
|
0.134192 |
0.134192 |
1
|
250
|
|
GO:0035223
|
leg disc pattern formation
|
0.138462 |
0.138462 |
1
|
18
|
|
GO:0009891
|
positive regulation of biosynthetic process
|
0.146409 |
0.146409 |
1
|
160
|
|
GO:0031328
|
positive regulation of cellular biosynthetic process
|
0.149561 |
0.149561 |
1
|
159
|
|
GO:0051716
|
cellular response to stimulus
|
0.150190 |
0.150190 |
1
|
374
|
|
GO:0010557
|
positive regulation of macromolecule biosynthetic process
|
0.151735 |
0.151735 |
1
|
131
|
|
GO:0016273
|
arginine N-methyltransferase activity
|
0.155172 |
0.155172 |
1
|
9
|
|
GO:0048522
|
positive regulation of cellular process
|
0.160427 |
0.160427 |
1
|
372
|
|
GO:0060173
|
limb development
|
0.164336 |
0.164336 |
1
|
47
|
|
GO:0035108
|
limb morphogenesis
|
0.166078 |
0.166078 |
1
|
47
|
|
GO:0035107
|
appendage morphogenesis
|
0.184365 |
0.184365 |
1
|
283
|
|
GO:0048519
|
negative regulation of biological process
|
0.186553 |
0.186553 |
1
|
706
|
|
GO:0009791
|
post-embryonic development
|
0.189538 |
0.189538 |
1
|
500
|
|
GO:0009892
|
negative regulation of metabolic process
|
0.193590 |
0.193590 |
1
|
372
|
|
GO:0007449
|
proximal/distal pattern formation, imaginal disc
|
0.196262 |
0.196262 |
1
|
21
|
|
GO:0007389
|
pattern specification process
|
0.203563 |
0.203563 |
1
|
537
|
|
GO:0051172
|
negative regulation of nitrogen compound metabolic process
|
0.216720 |
0.216720 |
1
|
250
|
|
GO:0031324
|
negative regulation of cellular metabolic process
|
0.219618 |
0.219618 |
1
|
319
|
|
GO:0005829
|
cytosol
|
0.228127 |
0.228127 |
1
|
425
|
|
GO:0006259
|
DNA metabolic process
|
0.231715 |
0.231715 |
1
|
242
|
|
GO:0009987
|
cellular process
|
0.235914 |
0.235914 |
3
|
6560
|
|
GO:0009890
|
negative regulation of biosynthetic process
|
0.242656 |
0.242656 |
1
|
282
|
|
GO:0016481
|
negative regulation of transcription
|
0.243724 |
0.243724 |
1
|
233
|
|
GO:0045934
|
negative regulation of nucleobase, nucleoside, nucleotide and nucleic acid metabolic process
|
0.252452 |
0.252452 |
1
|
250
|
|
GO:0031327
|
negative regulation of cellular biosynthetic process
|
0.253848 |
0.253848 |
1
|
282
|
|
GO:0009886
|
post-embryonic morphogenesis
|
0.259446 |
0.259446 |
1
|
412
|
|
GO:0044260
|
cellular macromolecule metabolic process
|
0.260978 |
0.260978 |
3
|
2878
|
|
GO:0048523
|
negative regulation of cellular process
|
0.262869 |
0.262869 |
1
|
635
|
|
GO:0043167
|
ion binding
|
0.265556 |
0.265556 |
1
|
1498
|
|
GO:0010605
|
negative regulation of macromolecule metabolic process
|
0.268230 |
0.268230 |
1
|
356
|
|
GO:0007552
|
metamorphosis
|
0.273794 |
0.273794 |
1
|
420
|
|
GO:0010629
|
negative regulation of gene expression
|
0.276112 |
0.276112 |
1
|
288
|
|
GO:0048569
|
post-embryonic organ development
|
0.276167 |
0.276167 |
1
|
343
|
|
GO:0016274
|
protein-arginine N-methyltransferase activity
|
0.281250 |
0.281250 |
1
|
9
|
|
GO:0030529
|
ribonucleoprotein complex
|
0.298647 |
0.298647 |
1
|
510
|
|
GO:0044085
|
cellular component biogenesis
|
0.299810 |
0.299810 |
1
|
632
|
|
GO:0043565
|
sequence-specific DNA binding
|
0.300971 |
0.300971 |
1
|
248
|
|
GO:0010558
|
negative regulation of macromolecule biosynthetic process
|
0.303014 |
0.303014 |
1
|
280
|
|
GO:0008276
|
protein methyltransferase activity
|
0.304762 |
0.304762 |
1
|
32
|
|
GO:0003824
|
catalytic activity
|
0.310936 |
0.310936 |
2
|
4106
|
|
GO:0008170
|
N-methyltransferase activity
|
0.314286 |
0.314286 |
1
|
33
|
|
GO:2000113
|
negative regulation of cellular macromolecule biosynthetic process
|
0.316071 |
0.316071 |
1
|
280
|
|
GO:0003676
|
nucleic acid binding
|
0.328842 |
0.328842 |
1
|
1855
|
|
GO:0043170
|
macromolecule metabolic process
|
0.329564 |
0.329564 |
3
|
3588
|
|
GO:0005575
|
cellular_component
|
0.330897 |
0.330897 |
3
|
8817
|
|
GO:0006281
|
DNA repair
|
0.331250 |
0.331250 |
1
|
106
|
|
GO:0005623
|
cell
|
0.346816 |
0.346816 |
3
|
6195
|
|
GO:0044464
|
cell part
|
0.346816 |
0.346816 |
3
|
6195
|
|
GO:0043226
|
organelle
|
0.353651 |
0.353651 |
2
|
3537
|
|
GO:0044424
|
intracellular part
|
0.354324 |
0.354324 |
3
|
4384
|
|
GO:0009058
|
biosynthetic process
|
0.355414 |
0.355414 |
2
|
2090
|
|
GO:0006807
|
nitrogen compound metabolic process
|
0.356248 |
0.356248 |
2
|
2093
|
|
GO:0009059
|
macromolecule biosynthetic process
|
0.360119 |
0.360119 |
2
|
1623
|
|
GO:0009887
|
organ morphogenesis
|
0.374384 |
0.374384 |
1
|
684
|
|
GO:0006139
|
nucleobase, nucleoside, nucleotide and nucleic acid metabolic process
|
0.374802 |
0.374802 |
2
|
1789
|
|
GO:0016835
|
carbon-oxygen lyase activity
|
0.375000 |
0.375000 |
1
|
54
|
|
GO:0007444
|
imaginal disc development
|
0.376006 |
0.376006 |
1
|
467
|
|
GO:0050896
|
response to stimulus
|
0.376795 |
0.376795 |
1
|
1548
|
|
GO:0044281
|
small molecule metabolic process
|
0.378352 |
0.378352 |
1
|
761
|
|
GO:0006950
|
response to stress
|
0.390827 |
0.390827 |
1
|
605
|
|
GO:0022625
|
cytosolic large ribosomal subunit
|
0.391892 |
0.391892 |
1
|
58
|
|
GO:0033554
|
cellular response to stress
|
0.397122 |
0.397122 |
1
|
276
|
|
GO:0003735
|
structural constituent of ribosome
|
0.398693 |
0.398693 |
1
|
183
|
|
GO:0005634
|
nucleus
|
0.408695 |
0.408695 |
2
|
1826
|
|
GO:0044267
|
cellular protein metabolic process
|
0.410493 |
0.410493 |
2
|
1505
|
|
GO:0048563
|
post-embryonic organ morphogenesis
|
0.415167 |
0.415167 |
1
|
323
|
|
GO:0022626
|
cytosolic ribosome
|
0.434783 |
0.434783 |
1
|
100
|
|
GO:0016740
|
transferase activity
|
0.438824 |
0.438824 |
1
|
1030
|
|
GO:0051252
|
regulation of RNA metabolic process
|
0.440308 |
0.440308 |
1
|
686
|
|
GO:0003677
|
DNA binding
|
0.444205 |
0.444205 |
1
|
824
|
|
GO:0005622
|
intracellular
|
0.446165 |
0.446165 |
3
|
4734
|
|
GO:0019538
|
protein metabolic process
|
0.463198 |
0.463198 |
2
|
2053
|
|
GO:0071840
|
cellular component organization or biogenesis
|
0.485282 |
0.485282 |
1
|
2108
|
|
GO:0043412
|
macromolecule modification
|
0.491525 |
0.491525 |
1
|
724
|
|
GO:0034645
|
cellular macromolecule biosynthetic process
|
0.495389 |
0.495389 |
2
|
1617
|
|
GO:0032774
|
RNA biosynthetic process
|
0.496008 |
0.496008 |
1
|
703
|
|
GO:0034641
|
cellular nitrogen compound metabolic process
|
0.497519 |
0.497519 |
2
|
1959
|
|
GO:0019222
|
regulation of metabolic process
|
0.507779 |
0.507779 |
1
|
1242
|
|
GO:0050789
|
regulation of biological process
|
0.509927 |
0.509927 |
1
|
2246
|
|
GO:0006412
|
translation
|
0.514942 |
0.514942 |
1
|
600
|
|
GO:0044249
|
cellular biosynthetic process
|
0.540182 |
0.540182 |
2
|
2034
|
|
GO:0010467
|
gene expression
|
0.547191 |
0.547191 |
2
|
1907
|
|
GO:0048513
|
organ development
|
0.548344 |
0.548344 |
1
|
1242
|
|
GO:0044238
|
primary metabolic process
|
0.551267 |
0.551267 |
3
|
4259
|
|
GO:0008757
|
S-adenosylmethionine-dependent methyltransferase activity
|
0.552381 |
0.552381 |
1
|
58
|
|
GO:0080090
|
regulation of primary metabolic process
|
0.556274 |
0.556274 |
1
|
1039
|
|
GO:0071841
|
cellular component organization or biogenesis at cellular level
|
0.556542 |
0.556542 |
1
|
1562
|
|
GO:0065007
|
biological regulation
|
0.561089 |
0.561089 |
1
|
2548
|
|
GO:0005737
|
cytoplasm
|
0.563502 |
0.563502 |
2
|
2378
|
|
GO:0032991
|
macromolecular complex
|
0.567899 |
0.567899 |
1
|
2151
|
|
GO:0031323
|
regulation of cellular metabolic process
|
0.569226 |
0.569226 |
1
|
1110
|
|
GO:0044422
|
organelle part
|
0.572743 |
0.572743 |
1
|
2176
|
|
GO:0043228
|
non-membrane-bounded organelle
|
0.573530 |
0.573530 |
1
|
1227
|
|
GO:0043232
|
intracellular non-membrane-bounded organelle
|
0.574428 |
0.574428 |
1
|
1227
|
|
GO:0032502
|
developmental process
|
0.575616 |
0.575616 |
1
|
2638
|
|
GO:0007560
|
imaginal disc morphogenesis
|
0.575758 |
0.575758 |
1
|
323
|
|
GO:0008150
|
biological_process
|
0.577615 |
0.577615 |
3
|
10616
|
|
GO:0009653
|
anatomical structure morphogenesis
|
0.581501 |
0.581501 |
1
|
1534
|
|
GO:0009889
|
regulation of biosynthetic process
|
0.599142 |
0.599142 |
1
|
888
|
|
GO:0051171
|
regulation of nitrogen compound metabolic process
|
0.617799 |
0.617799 |
1
|
905
|
|
GO:0006350
|
transcription
|
0.625331 |
0.625331 |
1
|
859
|
|
GO:0042254
|
ribosome biogenesis
|
0.626374 |
0.626374 |
1
|
57
|
|
GO:0060255
|
regulation of macromolecule metabolic process
|
0.631568 |
0.631568 |
1
|
1056
|
|
GO:0031326
|
regulation of cellular biosynthetic process
|
0.635595 |
0.635595 |
1
|
888
|
|
GO:0035120
|
post-embryonic appendage morphogenesis
|
0.638095 |
0.638095 |
1
|
268
|
|
GO:0043227
|
membrane-bounded organelle
|
0.653325 |
0.653325 |
2
|
2859
|
|
GO:0043231
|
intracellular membrane-bounded organelle
|
0.653801 |
0.653801 |
2
|
2856
|
|
GO:0003674
|
molecular_function
|
0.653878 |
0.653878 |
3
|
11064
|
|
GO:0048731
|
system development
|
0.657619 |
0.657619 |
1
|
1696
|
|
GO:0007275
|
multicellular organismal development
|
0.664929 |
0.664929 |
1
|
2296
|
|
GO:0050794
|
regulation of cellular process
|
0.666348 |
0.666348 |
1
|
2034
|
|
GO:0032501
|
multicellular organismal process
|
0.674757 |
0.674757 |
1
|
3315
|
|
GO:0006479
|
protein methylation
|
0.682927 |
0.682927 |
1
|
28
|
|
GO:0006355
|
regulation of transcription, DNA-dependent
|
0.684327 |
0.684327 |
1
|
620
|
|
GO:0045449
|
regulation of transcription
|
0.687166 |
0.687166 |
1
|
771
|
|
GO:0006464
|
protein modification process
|
0.689917 |
0.689917 |
1
|
684
|
|
GO:0007479
|
leg disc proximal/distal pattern formation
|
0.695652 |
0.695652 |
1
|
16
|
|
GO:0019219
|
regulation of nucleobase, nucleoside, nucleotide and nucleic acid metabolic process
|
0.699519 |
0.699519 |
1
|
903
|
|
GO:0010556
|
regulation of macromolecule biosynthetic process
|
0.705273 |
0.705273 |
1
|
850
|
|
GO:0006974
|
response to DNA damage stimulus
|
0.706522 |
0.706522 |
1
|
195
|
|
GO:0032259
|
methylation
|
0.728571 |
0.728571 |
1
|
51
|
|
GO:0007480
|
imaginal disc-derived leg morphogenesis
|
0.729167 |
0.729167 |
1
|
35
|
|
GO:0090304
|
nucleic acid metabolic process
|
0.736132 |
0.736132 |
2
|
1535
|
|
GO:0010468
|
regulation of gene expression
|
0.739954 |
0.739954 |
1
|
973
|
|
GO:2000112
|
regulation of cellular macromolecule biosynthetic process
|
0.763561 |
0.763561 |
1
|
850
|
|
GO:0008270
|
zinc ion binding
|
0.781440 |
0.781440 |
1
|
901
|
|
GO:0046914
|
transition metal ion binding
|
0.795721 |
0.795721 |
1
|
1153
|
|
GO:0016070
|
RNA metabolic process
|
0.798742 |
0.798742 |
1
|
1191
|
|
GO:0044444
|
cytoplasmic part
|
0.809942 |
0.809942 |
1
|
1863
|
|
GO:0006351
|
transcription, DNA-dependent
|
0.814170 |
0.814170 |
1
|
701
|
|
GO:0048707
|
instar larval or pupal morphogenesis
|
0.842767 |
0.842767 |
1
|
402
|
|
GO:0048856
|
anatomical structure development
|
0.858605 |
0.858605 |
1
|
2265
|
|
GO:0044446
|
intracellular organelle part
|
0.869797 |
0.869797 |
1
|
2162
|
|
GO:0005488
|
binding
|
0.882277 |
0.882277 |
1
|
5641
|
|
GO:0043229
|
intracellular organelle
|
0.899971 |
0.899971 |
2
|
3530
|
|
GO:0003002
|
regionalization
|
0.942272 |
0.942272 |
1
|
506
|
|
GO:0008168
|
methyltransferase activity
|
0.945946 |
0.945946 |
1
|
105
|
|
GO:0022613
|
ribonucleoprotein complex biogenesis
|
0.947917 |
0.947917 |
1
|
91
|
|
GO:0002165
|
instar larval or pupal development
|
0.954000 |
0.954000 |
1
|
477
|
|
GO:0046872
|
metal ion binding
|
0.967936 |
0.967936 |
1
|
1449
|
|
GO:0035114
|
imaginal disc-derived appendage morphogenesis
|
0.975524 |
0.975524 |
1
|
279
|
|
GO:0035110
|
leg morphogenesis
|
0.978723 |
0.978723 |
1
|
46
|
|
GO:0048737
|
imaginal disc-derived appendage development
|
0.986014 |
0.986014 |
1
|
282
|
|
GO:0043169
|
cation binding
|
0.999332 |
0.999332 |
1
|
1497
|
|
GO:0000000
|
root
|
1.00000 |
1.00000 |
3
|
12747
|
|
GO:0003700
|
sequence-specific DNA binding transcription factor activity
|
1.00000 |
1.00000 |
1
|
395
|